Mass Spectrometry and Analytical Pharmacology Shared Resource (MSAPSR)
Metabolomics Instrumentation and Services
Established in 2008, the mission of the Metabolomics component of the Mass Spectrometry and Analytical Pharmacology Shared Resource (MSAPR) is to provide an in-house resource to facilitate cutting-edge basic science, clinical and translational research studies among LCCC members and GU researchers. The Metabolomics component of the MSAPSR provides mass spectrometry based high quality laboratory and data analysis services to researchers including comprehensive metabolomics services such as global profiling as well as multiple reaction monitoring based targeted quantitation of metabolites, lipids and drugs. These services are applicable to an array of matrices including urine, blood, tissue, cells, cerebrospinal fluid among others. The Metabolomics component of the MSAPSR is directed by Dr. Amrita Cheema assisted by Dr. Shivani Bansal who serves as the facility manager. The MSAPSR is available to researchers for routine and specialized metabolomics applications. The aim of the core is to provide an “in house” resource, rich in technical expertise, that fosters the free flow of experimental knowledge and collaborations across Georgetown University and the LCCC Research Consortium.
Metabolomics Services offered by the Shared Resource include:
- Advice and guidance on the use and applications of methodologies and on experimental design and data interpretation.
- Provide technical support for developing grant proposals and scientific manuscripts.
- Research and development to fit users’ specific needs. New assays can be tailored according to the customer request.
- Training on post-processing of LC-MS data
- Mentoring of technicians and students involved in projects via online and onsite training sessions
Metabolomics Technologies offered by the Shared Resource include:
- Untargeted profiling – UPLC-QToF-MS
- Targeted assays (UPLC-MRM-MS): Multiple reaction monitoring in conjunction with ultra-performance chromatography is employed for the analysis of different sample matrices including cell extracts, plasma, serum, urine, fecal, and tissue. The standard assays that are available to researchers include:
- Absolute IDQ p180 Kit – Biocrates
- Amino acid panel
- Lipid mediator panel
- Oxylipin panel
- Biogenic amines
- Bile acid
- TCA cycle metabolites
- Metabolites in single carbon metabolism
- Vitamins
- Nucleoties/Nucleosides
- Sphingolipids
- Phosphocholine
- Carnitines
- Cardiolipins
- 2,000 deep lipidomics panel
- Polar metabolites panel for 350 endogenous small molecules
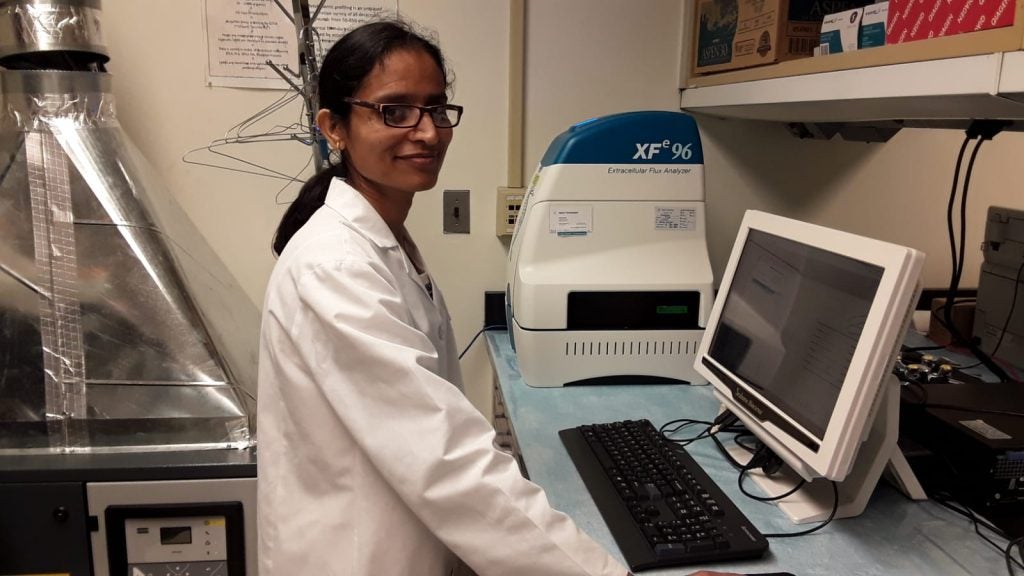
- Mitochondrial respiration and glycolysis in live cells using the Seahorse Extracellular Flux Analyzer
- Nuclear magnetic resonance-based analysis
Metabolomics Instrumentation
Waters Xevo G3 Q-TOF: The core facility is equipped with the G3 QToF online with an ultra-performance liquid chromatography (UPLC) system known for its exceptional analytical capabilities. This instrument utilizes advanced chromatographic techniques to separate, identify, and quantify components in complex mixtures for global metabolomic profiling analyses.
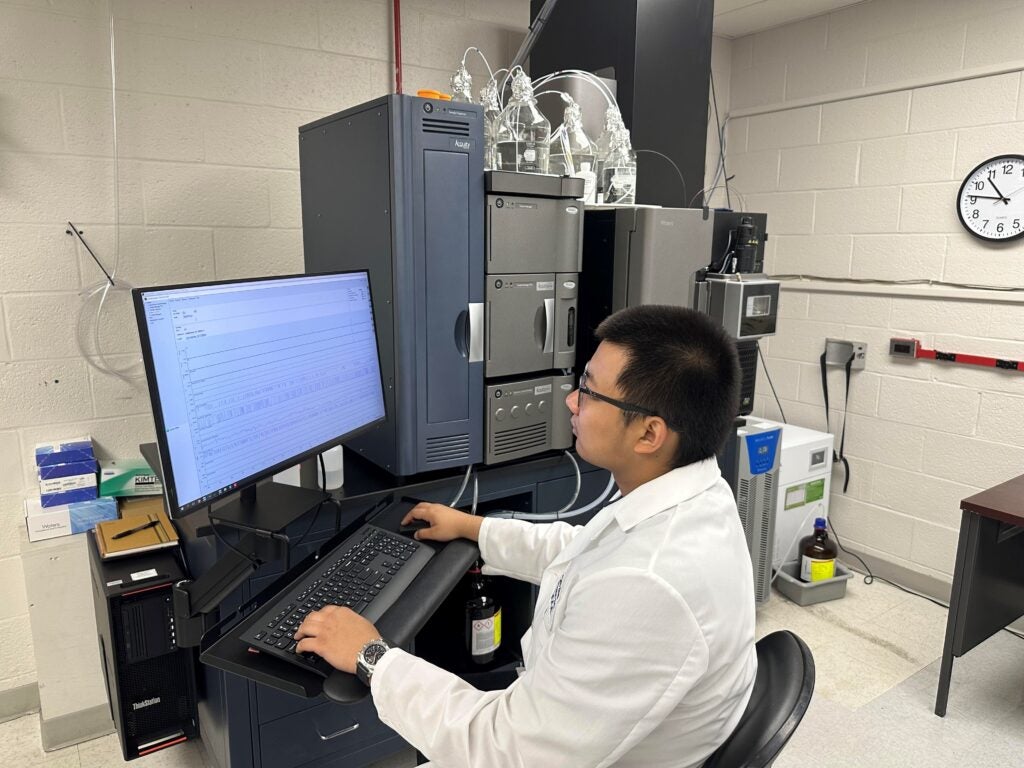

Bruker timsTOF Pro 2: The core facility houses the Bruker TimsTOF Pro2, a state-of-the-art mass spectrometer for high sensitivity workflow in metabolomics and lipidomics. This instrument utilizes a unique combination of trapped ion mobility spectrometry (TIMS) and time-of-flight (TOF) technologies, providing high-resolution, high-sensitivity for analysis of complex biological samples.
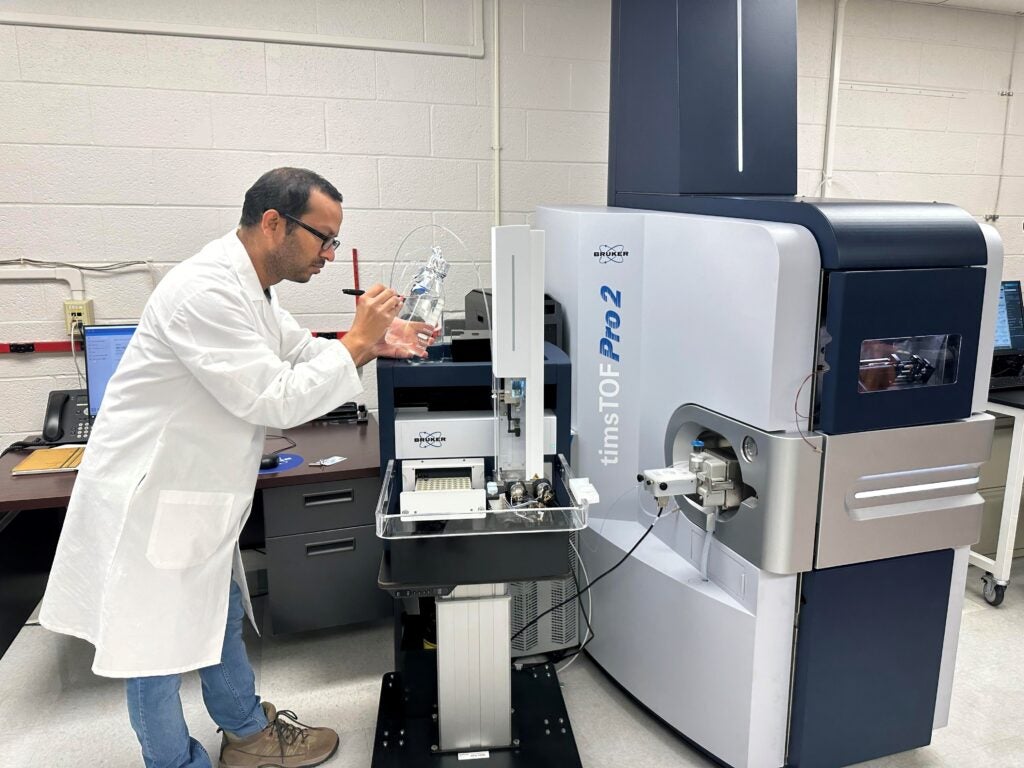

SCIEX 7500 Qtrap: The metabolomics core facility houses the SCIEX 7500 Qtrap, a hybrid triple quadrupole/linear ion trap technology, that supports multiplexed quantitation of analytes of interest. The SCIEX 7500 Qtrap is routinely used in the core for multiple reaction monitoring based quantitation for metabolomics and lipidomics studies.

Agilent HPLC with fraction collector: The core facility is equipped with an Agilent High-Performance Liquid Chromatography (HPLC) system with a fraction collector, a crucial instrument for enhancing metabolome and lipidome coverage. HPLC is a technique used to separate, identify, and quantify each component in a mixture. The Agilent HPLC system in our facility allows for offline fractionation of analytes, a process that separates the components of a sample based on their chemical properties. This separation technique increases the coverage of the metabolome and lipidome, providing a more comprehensive analysis of the sample. The fraction collector component of the system further enhances this process by collecting the separated fractions for further analysis.

Waters Xevo TQ-S: The Xevo TQ-S is a triple quadrupole system that supports MRM-MS based targeted analysis for kit-based assays, biomarker validation and pharmacokinetics studies.

Varian Nuclear Magnetic Resonance (NMR): The Varian Nuclear Magnetic Resonance (NMR) housed in the metabolomics is a powerful analytical tool that provides detailed information about the physical and chemical properties of chemical compounds. NMR spectroscopy exploits the magnetic properties of certain atomic nuclei to provide detailed molecular information.

SCIEX 5500 Qtrap: MSR also houses the 5500 QTrap equipped with differential mobility spectrometry (Lipidyzer) to support reliable and robust quantitative analysis of complex lipid panels. This instrument supports an array of research projects focused on lipidomics research, enabling scientists to gain a comprehensive understanding of lipid metabolism and its role in various biological processes and diseases. In addition to its lipid profiling capabilities, the 5500 QTrap also supports routine targeted workflows, making it a versatile tool for a wide range of research applications.

Leco GC-ToF: The core facility is equipped with the Leco GC-ToF, a high-performance gas chromatography time-of-flight mass spectrometer used for untargeted profiling of a wide range of matrices, including cell extracts, cell media, plasma, serum, urine, fecal samples, and tissue.

Sea Horse XF Analyzer: Enables simultaneous measurement of mitochondrial respiration and glycolysis in live cells in real time. This powerful tool provides researchers with a comprehensive understanding of cellular metabolic function, offering insights into the role of cellular metabolism in health and disease.